Research Article - (2018) Volume 8, Issue 5
Genetic Diagnosis in Children with Epilepsy and Developmental Delay/Mental Retardation Using Targeted Gene Panel Analysis
- Corresponding Author:
- Jyh-Feng Lu, PhD
School of Medicine, Fu Jen Catholic University No.510
Zhongzheng Rd. Xinzhuang Dist, New Taipei City 24205, Taiwan
Tel: 886-2-29053464
Fax: 886-2-29052096
Abstract
Purpose
Epileptic encephalopathies (EEs) are the severe form of childhood epilepsy and phenotypically heterogeneous disorders with different underlying genetic defects. EEs are always accompanied with developmental delay/mental retardation and behavior problems. Finding the genetic basis of epileptic encephalopathies with developmental delay/mental retardation can be valuable not only for diagnosis but also for guiding treatment and providing disease prognosis.
Methods and Results
A customized panel of 90 genes related to epileptic encephalopathies was utilized to screen for potential genetic variants via targeted next generation sequencing (NGS). A total of 78 children with epilepsy and developmental delay/mental retardation were analyzed with an average read depth of 265.3 ± 68.3X. Mutations were found in 23 (9 boys and 14 girls) probands, and the overall mutation identification rate was 29.5% (25.0% for males and 33.3% for females). Fifteen (65.2%) mutations involve ion channels, including SCN1A, KCNT1, SCN2A, SCN8, KCNB1 and KCNQ2. The other (34.8%) mutations involve CDKL5, ALG13, GFAP, SNAP25, STXBP1 and SYNGAP1. Sixteen (69.6%) of the identified mutations were confirmed to be de novo and one (4.3%) was found to be paternal mosaicism. Channelopathies were found to be the major cause of both early onset (≤6 months) and later onset of unclassified EE as well as febrile seizure cases. On the contrary, patients with infantile spasms or West syndrome were mainly caused by mutations in non-ion channel proteins, whereas patients of epilepsy with developmental delays and/or mental retardation were caused by mutations in both channel and non-channel genes.
Conclusion
NGS is a valuable and reliable diagnostic tool to detect the gene mutation in epileptic children with developmental delay/mental retardation. This cost-effective method shortens the course from seizure onset to genetic diagnosis. By understanding the gene mutations and genotypephenotype correlation better, clinical practitioners could provide the optimal anticonvulsant and treatment.
Keywords
Epileptic encephalopathy (EE), Developmental delay (DD), Febrile seizure (FS), Infantile spasms (IS), West syndrome (WS), Targeted next generation sequencing (NGS), Mutation
Introduction
Epilepsy is a common neurological disorder in children with broad phenotypic spectrum. Epileptic encephalopathies (EEs) are the most severe form of epilepsy. Heterogeneous clinical manifestations characterized by refractory seizures and developmental delay, which are often accompanied with various psychiatric comorbidities [1,2], age of onset, seizure types, interictal condition, electroencephalography (EEG) pattern and causes are variable in different epileptic encephalopathies. About one third of all EE cases are referred to as cryptogenic cases, where genetic factors are considered to play an important role [3-5]. More than 100 genes have been found to be involved in patients with epileptic encephalopathies [6]. Next generation sequencing (NGS) techniques, which allow massive parallel sequencing of as many genes as desired, are fast and cost-efficient diagnostic tools used to analyze the genetic cause of epileptic encephalopathies [6]. The genetic diagnostic yield can be increased from less than 10% to 28% in patients with epileptic encephalopathy using targeted NGS panels [7]. Not only the genes encoding membrane ion channel proteins, but also genes encoding proteins other than ion channels are now being identified as the cause of EE by NGS [8]. The present study is a retrospective cohort study for genetic testing in children with cryptogenic epilepsy and developmental delay/ mental retardation (DD/MR) using NGS to identify the genetic characteristics and correlation with clinical features.
Materials and Methods
▪ Participants
Seventy-eight Taiwanese children (36 boys and 42 girls) with epilepsy and DD/MR were collected with informed consent according to protocol (105142F) approved by the Internal Review Board of Far Eastern Memorial Hospital. Inclusion criteria were 70 children aged 7 months to 18 years 5 months (average 6 years 8 month ± 4 years 6 months) and 8 long-term follow-up patients (aged 20 years 4 months to 26 years 8 months, average 23 years ± 2 years 4 months) with cryptogenic epileptic encephalopathy. Children with family history of known neurological disorders, possible damage or infection of central nervous system (CNS), and known metabolic disturbance were excluded. Twenty three (13 males and 10 females) had early onset epileptic encephalopathy (EoEE) with seizure onset before 6 month-old with or without suppression-burst on EEG. Ten (3 males and 7 females) had unclassified epileptic encephalopathy (EE) with seizure onset after 6-month-old. Twenty (6 males and 14 females) exhibited either infantile spasms (IS) or West syndrome (WS). Twenty one (11 males and 10 females) presented with epilepsy with developmental delay and/or mental retardation (E-DD/MR). Four (3 males and 1 females) had febrile seizure (FS) with developmental delay and/or mental retardation (FS-DD/ MR). Genomic DNA (gDNA) were extracted from peripheral white blood cells and were evaluated both quantitatively and qualitatively by an Invitrogen™ Qubit™ 3.0 Fluorometer (ThermoFisher Scientific Inc., Waltham, MA, USA) and an Agilent 2200 TapeStation system (Agilent Technologies, Santa Clara, CA, USA) before being used for NGS analysis.
▪ Library preparation and sequencing
A customized panel of 90 genes related to epileptic encephalopathy was created using SureDesign (Agilent Technologies, Santa Clara, CA, USA) and used as the HaloPlex probe capture library for the enrichment of DNA targets for mutational screening. For the generation of standard target-enriched libraries, the Agilent HaloPlex Target Enrichment System kit was used following the protocol for Illumina Hiseq paired-end sequencing library (Agilent Technologies, Santa Clara, CA, USA). The gDNA was digested in eight different restriction reactions (225 ng DNA/reaction), each containing two restriction enzymes. All restriction reactions were validated by using High Sensitivity D1000 ScreenTape in an Agilent 2200 TapeStation (Agilent Technologies). The collection of gDNA restriction fragments was hybridized to the HaloPlex probe capture library at 54°C for 3 hours for target enrichment. The circularized target DNA-HaloPlex probe hybrids were captured on streptavidin beads (HaloPlex Magnetic Beads, Agilent Technologies) and DNA ligase was then added to close nicks in the hybrids. Elution of captured DNA was carried out using NaOH, and 20 uL of the eluted supernatant was used as template for PCR amplification. Twenty PCR cycles were performed to amplify target libraries and the PCR amplicons were purified using Agencourt AMPure XP beads (Beckman Coulter, Brea, CA, USA) for NGS. DNA sequencing was performed on an Illumina Miseq system using 150PE protocol (Illumina, Inc. San Diego, CA, USA).
▪ Variants analysis
The sequencing data obtained from Illumina Miseq instrument were subjected to QC verification and the qualified read data went through genomic alignment against Ensembl human reference genome database using BWA software [9]. Variant calling of aligned sequences were created by Genome Analysis Toolkit (GATK) [10] and further analyzed using Illumina VariantStudio 3.0 software (Illumina, Inc.). Identified variants in probands and their parents were subjected to Sanger sequencing for further validation.
Results
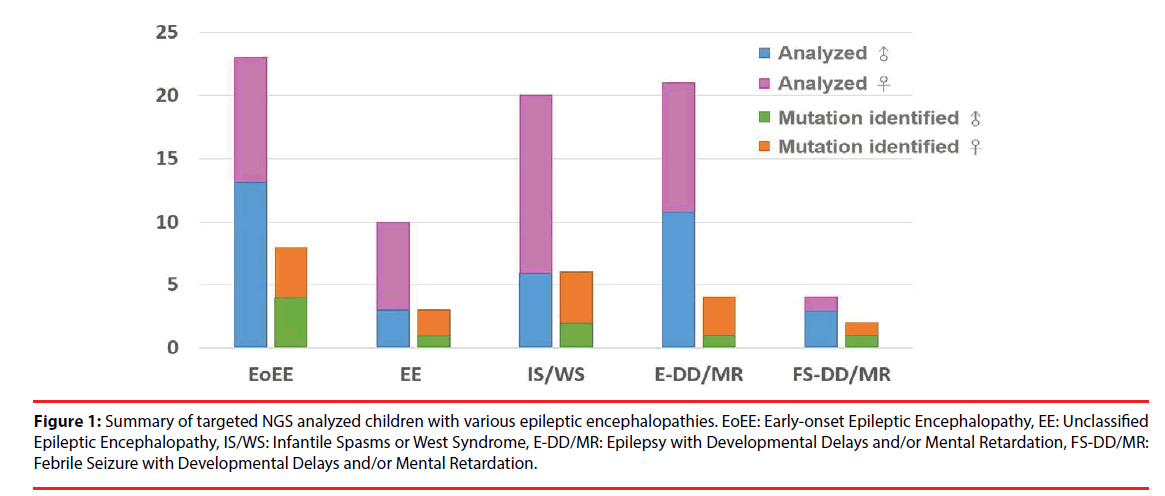
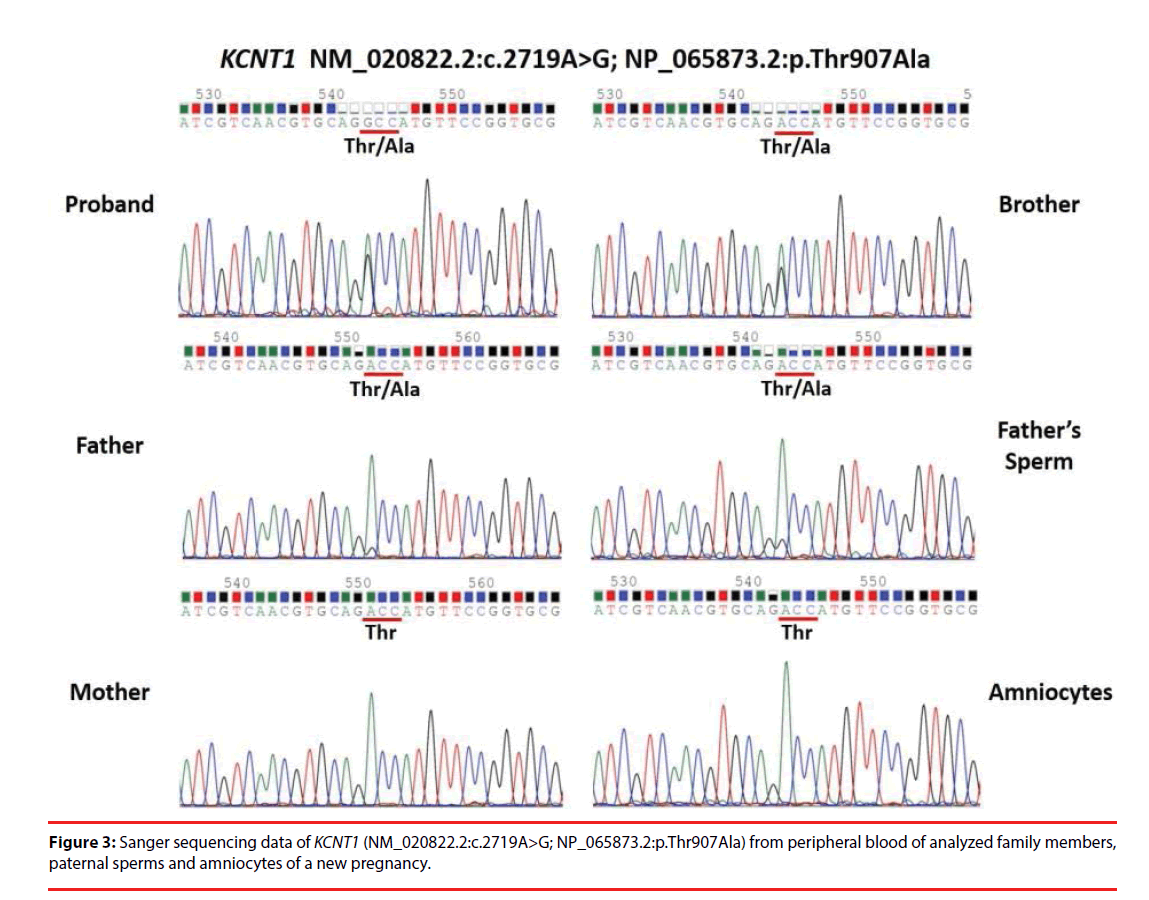
A total of 78 Taiwanese patients with epileptic encephalopathies were collected and analyzed using targeted NGS. The average read depth on target sequences was 265.3 ± 68.3X (ranged from 122-464X). As shown in Figure 1, mutations were found in 23 (9 boys and 14 girls) probands (Supplementary materials); the overall mutation identification rate via this designed panel was 29.5% (25.0% for males and 33.3% for females). Among them, 8 individuals were EoEE (identification rate: 8/23, 34.8%), 6 were IS/WS (6/20; 30.0%), 4 were E-DD/MR (4/21, 19.0%), 3 were EE (3/10, 30.0%), and 2 were FS-DD/MR (2/4, 50.0%). The details of patient clinical presentations and identified mutations are listed in Table 1. Fifteen (65.2%) mutations involve ion channels, including 5 (21.7%) in SCN1A, 4 (17.4%) in KCNT1, 2 (8.7%) each in SCN2A and SCN8A, and one (4.3%) of each for KCNB1 and KCNQ1. The other 8 (34.8%) mutations include 3 (13.0%) in CDKL5 and one (4.3%) of each for ALG13, GFAP, SNAP25, STXBP1 and SYNGAP1 (Figure 2). Sixteen (69.6%) of the identified mutations were confirmed to de novo and one (4.3%) was found to be paternal mosaicism (Figure 3). Confirmation of paternity for de novo cases was performed by genotyping a short tandem repeat (STR) marker. Seven de novo mutations were identified in patients with EoEE (7/23, 30.4%), 5 for IS/WS (5/20, 25.0%), 1 (1/10, 10.0%) for EE, and 3 (3/21, 14.3%) for E-DD/MR. De novo mutation rate was 55.6% (5/9) for males and was 78.6% (11/14) for females. The others include 2 familial mutations (8.7%) and 5 (21.7%) were not determined due to the lack of parental samples. Seizure onset to diagnosis among 23 individuals with mutation identified by targeted NGS ranges from 1 month to 21 years and 8 months (average 6 years and 7 months ± 5 years and 3 months). The frequency of all identified mutant alleles were relatively rare among all populations, including Asians, and the highest frequency reported was 8 x 10-6 for rs267607518 of the GFAP in the Exome Aggregation Consortium (ExAC) database. Twelve (12/23, 52.2%) of the 23 identified mutations have never been reported previously.
Figure 1: Summary of targeted NGS analyzed children with various epileptic encephalopathies. EoEE: Early-onset Epileptic Encephalopathy, EE: Unclassified Epileptic Encephalopathy, IS/WS: Infantile Spasms or West Syndrome, E-DD/MR: Epilepsy with Developmental Delays and/or Mental Retardation, FS-DD/MR: Febrile Seizure with Developmental Delays and/or Mental Retardation.
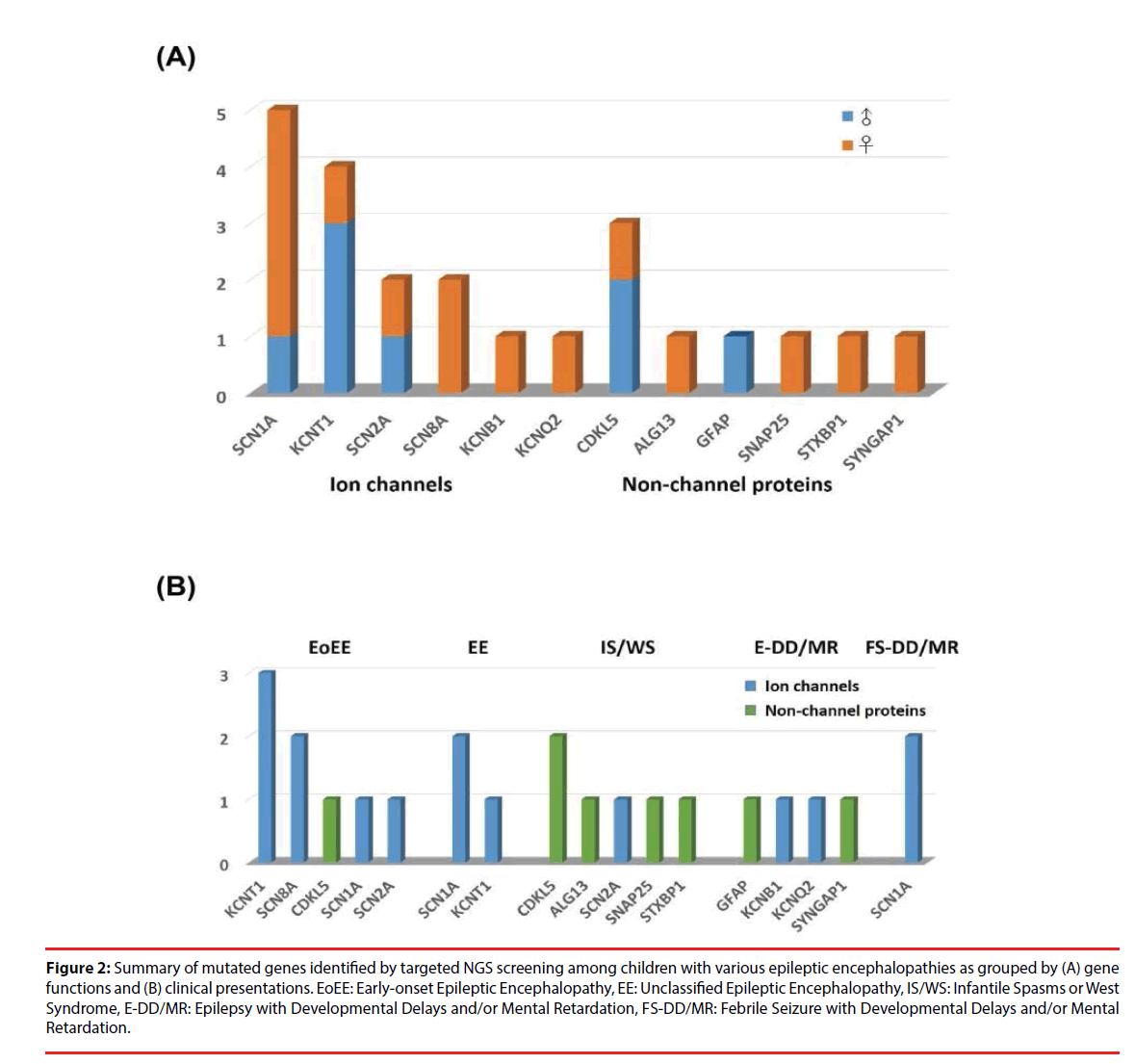
Figure 2: Summary of mutated genes identified by targeted NGS screening among children with various epileptic encephalopathies as grouped by (A) gene functions and (B) clinical presentations. EoEE: Early-onset Epileptic Encephalopathy, EE: Unclassified Epileptic Encephalopathy, IS/WS: Infantile Spasms or West Syndrome, E-DD/MR: Epilepsy with Developmental Delays and/or Mental Retardation, FS-DD/MR: Febrile Seizure with Developmental Delays and/or Mental Retardation.
| Gene (GRCh37 coordinate) | Sex | Age of Diagnosis | Seizures (Onset age) | Mutation | Parental Origin/ Novel(CS) | Diagnosis | ||
|---|---|---|---|---|---|---|---|---|
| SCN1A (NM_001202435.1) | F | 4y8m | FS, GS, PS | c.181delC | De novo | EoEE, MR | ||
| (chr2:166,929,951) | (4m) | p.L61Ffs*31 | Novel | |||||
| F | 2y5m | FS, GS, PS | c.482delT | NA | Complex FS, DD | |||
| (chr2:166,911,268) | (4m) | p.F161Sfs*11 | Novel | |||||
| F | 23y2m | FS, GS, PS | c.1134_1136delAAT | non-M | EE, DD/MR | |||
| (chr2:166,904,171) | (1y) | p.M379del | Novel | |||||
| F | 8y | FS,GS,PS | c.1177C > T | De novo | EE, DD/MR | |||
| (chr2:166,903,480) | (1y) | p.R393C | rs398123588(P) | |||||
| M | 9y | FS,GS,PS | c.4555C > A | Paternal | GEFS+, DD, ADHD | |||
| (chr2:166,852,549) | (1y1m) | p.P1519T | rs796053021(U)*,# | |||||
| KCNT1 (NM_020822.2) | M | 1y8m | SE | c.1041G > C | De novo | EoEE, Expired | ||
| (chr9:138,656,882) | (1m) | p.E347D | Novel*,# | |||||
| F | 8y5m | GS, PS, TS | c.1283G > A | NA | EoEE, DD/MR | |||
| (chr9:138,657,552) | (2m) | p.R428Q | rs397515402(P) | |||||
| M | 8y8m | GS,PS | c.785G > A | non-M | EE, DD/MR | |||
| (chr9:138,650,285) | (1y) | p.R262Q | SCV000589778.2(LP)*,# | |||||
| M | 1y7m | GS,PS | c.2719A > G | De novo (Paternal mosaicism) | EoEE, DD/MR | |||
| (chr9:138,670,658) | (2m) | p.T907A | Novel*,# | |||||
| SCN2A (NM_001040142.1) | F | 11y | FS,IS GTCS, TS, MS | c.1270G > A | De novo | WS, Rett-like syndrome, DD/MR [42] | ||
| (chr2:166,170,505) | (8m) | p.V424M | Novel*,# | |||||
| M | 25d | MS, TS, PS | c.573G > T | De novo | EoEE, DD/MR | |||
| (chr2:166,165,272) | (3d) | p.W191C | Novel*,# | |||||
| SCN8A (NM_014191.3) | F | 2y7m | GS | c.2300C > T | De novo | EoEE, DD/MR | ||
| (chr12:52,145,307) | (1m) | p.T767I | rs797045013(P) | |||||
| F | 1y | PS,GS | c.4423G > A | De novo | EoEE, DD | |||
| (chr12:52,184,185) | (4m) | p.G1475R | rs796053216(C)# | |||||
| KCNB1 (NM_004975.2) | F | 7y4m | GS, PS | c.629C > T | De novo | Epilpesy, MR | ||
| (chr20:47,991,468) | (2y6m) | p.T210M | Novel*,# | |||||
| KCNQ2 (NM_172107.2) | F | 10y10m | GS,TS | c.1741C > T | Maternal | Epilepsy, DD | ||
| (chr20:62,044,825) | (7d) | p.R581* | rs118192236(P) | |||||
| CDKL5 (NM_003159.2) | M | 3y | IS, GS | c.1670C > G | NA | WS, MR | ||
| (chrX:18,622,714) | (2m) | p.S557* | Novel | |||||
| F | 3y2m | MS, TS | c.2635_2636delCT | De novo | EoEE, DD/MR, Autism | |||
| (chrX:18,646,629) | (2m) | p.L879Efs*30 | rs61753251(P) | |||||
| M | 9y1m | IS | c.849dupT | De novo | WS, DD/MR | |||
| (chrX:18,616,605) | (3m) | p.D284* | Novel | |||||
| ALG13 (NM_001099922.2) | F | 5y4m | IS | c.320A > G | De novo | WS, DD/ MR | ||
| (chrX:110,928,268) | (5m) | p.N107S | rs398122394(P) | |||||
| GFAP (NM_001242376.1) | M | 6y9m | PS | c.[218T > C; 259 G > T] | De novo | Epilepsy, DD/MR | ||
| (chr17:42,992,637; chr17:42,992,596) |
(8m) | p.[M73T; V87F] | rs61060395(P); rs267607518(P) |
|||||
| SNAP25 (NM_003081.3) | F | 5y10m | IS,TS | c.149T > C | De novo | IS, DD/ MR | ||
| (chr20:10,265,406) | (7m) | p.L50S | Novel*,# | |||||
| STXBP1 (NM_003165.3) | F | 6y11m | IS, TS, GS, PS | c.224A > G | De novo | WS, MR | ||
| (chr9:130,420,708) | (4m) | p.Y75C | rs796053352*,# | |||||
| SYNGAP1 (NM_006772.2) | F | 11y | GS | c.1786C > A | De novo | Epilepsy, DD/MR | ||
| (chr6:33,408,615) | (1y3m) | p.R596S | Novel*,# | |||||
Table 1: Summary of clinical presentations and identified mutations.
As shown in Figure 2B, channelopathy was the major cause of EoEE and EE. EoEE involves mainly KCNT1 followed by SCN8A, SCN1A and SCN2A as well as a non-channel protein CDKL5. EE involves SCN1A and KCNT1. In addition, two FS-DD/MR patients also carry mutations in the ion channel SCN1A. On the contrary, patients with IS/WS mainly resulted from mutations in non-ion channel proteins, including CDKL5, ALG13, SNAP25 and STXBP1 as well as one caused by SCN2A mutation. Patients with E-DD/MR were caused by mutations in both channel and non-channel proteins, including KCNB1 and KCNQ2, as well as GFAP, and SYNGAP1.
Discussion
It is often difficult to diagnose accurately for young children with seizures due to continuous evolving nature of age-dependent epileptic encephalopathies [11]. The phenotypic spectrum is broad for the same mutated gene and the phenotype may be similar among mutations in different genes. Therefore, a cost-effective genetic screening can help the practitioner make the diagnosis. A panel of 90 genes for targeted NGS screening was developed to facilitate the identification of underlying genetic causes in order to assist clinical diagnosis and prognosis as well as to administer therapeutic choices. Targeted NGS analysis with an average read depth of ~265X, mutations were found in 23 (29.5%) out of 78 children with epileptic encephalopathies and developmental delay/mental retardation. The average read depth among these 23 individuals was 264.5 ± 82.0X (ranged from 137-464X) as compared with that of 265.7 ± 62.5X (ranged from 122-402X) among the other 55 individuals without identified mutation; the difference is not statistically significant (p=0.944). Furthermore, 4 (17.4%) of 23 individuals with identified mutation had an average read depth of less than 200X (157.3 ± 14.7X, ranged from 137~170X). These results indicate that an average read depth above 250X is sufficient for this targeted panel to perform mutational screening in children with epileptic encephalopathies and DD/MR. Compared with two recent studies of about 18% of genetic diagnosis rate in patients with epilepsy [12,13], the higher mutation identification rate in our study would likely be due to the use of a higher NGS read depth (250X vs. 30X), and a more focused patient cohort (mainly EoEE and IS/WS vs. clinically referred epilepsy patients).
The majority (65.2%) of the genetic causes for Taiwanese children with epilepsy and DD/MR are due to channelopathy as previously suggested for idiopathic epilepsies [14]. Most common involvement are Na+ channels, including SCN1A, SCN2A and SCN8A representing the subunit 1 (Nav1.1), 2 (Nav1.2) and 8 (Nav1.6), respectively, of the neuronal voltage-gated Na+ channel; followed by defects in K+ channels, including KCNT1, KCNB1 and KCNQ2 representing sodium-activated K+ channel subfamily T member 1 (Slo2.2), voltage-gated K+ channel subfamily B member 1 (Kv2.1) and subfamily Q member 2 (Kv7.2), respectively. Mutations in SCN1A have been reported to be associated with generalized epilepsy with febrile seizures plus (GEFS+) type 2 as the major culprit (~75%) for Dravet syndrome (DS) [15]. Among 5 patients identified to carry SCN1A mutations, two each were originally classified as EE and FS-DD/MR in addition to one as EoEE. One patient with EE was later confirmed to be DS. Three patients also exhibit MR which is commonly observed in DS due to developmental plateauing [16]. The significant features of our patients with SCN1A mutation is fever sensitive seizure. Mutations in SCN2A were reported to be associated with epileptic syndromes ranging from benign familial neonatal-infantile seizures [17] to severe epileptic encephalopathies, such as Ohtahara syndrome, malignant migrating focal seizures of infancy (MMFSI) [18,19] and late onset ataxia and autism spectrum disorders [20,21]. Two missense mutations were identified among a WS and an EoEE patient; the former exhibit Rett-like features and the latter is confirmed to be MMFSI. The Mutations in SCN8A have been reported for early infantile and unclassified EE, as well as in patients with Dravet-like presentation [22,23]. Similar to those of SCN1A in DS and SCN2A in MMFSI, the majority of SCN8A mutations in EE cases resulted from de novo mutations [13,16,24]. Six (66.7%) out of 9 analyzed patients with mutations in Na+ channels are de novo, which are consistent with observations in prior studies.
KCNT1, KCNB1 and KCNQ2 are members of large and diverse families of K+ channels, the disease causing mutations were found to be associated respectively with MMFSI [25] and with EoEE [26] via whole exome sequencing. KCNT1 is responsible for hyperpolarization modulation [27] and KCNB1 is the main contributor to delayed rectifier K+ current that is critical for membrane repolarization following repetitive firing of action potentials [26]. Four missense mutations in KCNT1 were identified in 3 EoEE and 1 EE patients. Two of them, including the only EE patient, were confirmed to be MMFSI. Importantly, the genetic diagnosis of KCNT1 mutations helps the clinicians confirm the diagnosis of MMFSI. One missense mutation (R262Q) is localized to cytoplasmic end of the S4 transmembrane domain, the others are localized to cytoplasmic C-terminal domain, including E347D next to S6 transmembrane domain, R428Q in the first regulator of conductance of K+ (RCK) domain and T907A in the NAD+ binding domain. All identified mutations, except R428Q, are de novo. For KCNB1, a missense mutation (T210M) was localized to the extracellular linker between S1 and S2 segments of the transmembrane domain. KCNQ2 encodes for Kv7.2 K+ channel subunits underlying the neuronal voltage-gated M-current (IKM) [28]. A maternally inherited nonsense mutation was identified in a familiar case of E-DD/MR.
The rest (34.8%) of the mutations were found in genes encoding non-ion channel proteins, including 3 in CDKL5, and one in each for ALG13, GFAP, SNAP25, STXBP1 and SYNGAP1. CDKL5 encodes for cyclin dependent kinase like Ser/Thr protein kinase and is one of the most frequently involved gene for X-linked IS, WS and atypical Rett syndrome predominantly in females [29,30]. A nonsense mutation (S557*) and a single nucleotide duplication leading to premature termination (D284*) of CDKL5 were identified in two male IS patients. In addition, a recurrent two nucleotide deletion frameshift mutation (c.2635_2636delCT; p.L879Efs*30) was found in a female EoEE patient [31]. Only 25 mutations in CDKL5 were previously reported in boys, as compared with 131 reported in females. The fact that two out of six of our analyzed male IS patients carry mutations warrants the necessity of mutational analysis of CDKL5 in males with IS. Furthermore, two other analyzed female IS patients were found to carry de novo mutation in ALG13 and SNAP25 genes encoding a subunit of a bipartite UDPN- acetylglucosaminyltransferase involving in N-linked glycosylation in endoplasmic reticulum and the synaptosome associated protein 25 for the regulation of neurotransmitter release in presynaptic plasma membranes, respectively.
Mutations in STXBP1 have been reported to be associated with intellectual disability with or without epilepsy as well as EoEE mainly Ohtahara syndrome [32]. Some (~75%) patients with Ohtahara syndrome are known to develop WS between 2~6 months of age and could further evolve into Lennox-Gastaut syndrome (LGS) [33,34]. The gene product of STXBP1, syntaxin binding protein 1, is involved in modulating the release of synaptic vesicles via interaction with syntaxin A and soluble N-ethylmaleimide-sensitive factor attachment protein receptors (SNAREs) complex [35]. A de novo missense mutation was found in the STXBP1 of a female Taiwanese WS patient. In addition, a de novo missense mutation in SYNGAP1 was also identified in a female E-DD/ MR patient. SYNGAP1 encodes for synaptic Ras GTPase activating protein 1, which is highly enriched at excitatory synapses and is important for proper neuronal development and glutamate receptor trafficking [36]. It is previously reported to be associated with intellectual disability (ID), EE or autism spectrum disorders (ASD) [37,38]. Thus, this supports the importance of genes involving trafficking of synaptic vesicles and its regulation in the pathogenesis of epileptic encephalopathies.
Double heterozygous missense mutations (c. [218T>C; 259 G>T]) in GFAP were identified in a boy with EoEE, who is confirmed to be diagnosed with Alexander disease (AD). Both mutations were found to be transmitted from the father, most likely due to de novo germline mutation [39]. GFAP is located on chromosome 17q21 and encodes glial fibrillary acidic protein which is the major intermediate filament proteins of mature astrocytes and is reported to be associated with a rare autosomal dominant neurological disorder AD [40]. Both missense mutations were localized to the 1st α-helical coil of the 4 coiled subdomain within the central rod domain of GFAP near known mutation hotspots (R79 and R239) [41].
In summary, overall mutation identification rate was 29.5% (23/78) among Taiwanese patients using targeted gene panel for the investigation of underlying genetic causes of epilepsy with DD/ MR. The diagnostic yield is highest in the EoEE subgroup. The majority of identified mutations involve genes encoding ion channels (65.2%) and more than two-thirds of identified mutations (69.6%) are de novo. Traditionally, the diagnosis of epileptic encephalopathies is based on clinical manifestations and EEG findings. Advances in high-throughput DNA sequencing technology have greatly facilitated detection of underlying genetic culprits of patient with various epileptic syndromes. Based on this study, targeted gene panels for epilepsy with DD/MR could facilitate the identification of mutation within a month from the onset of seizure. Thus, it would be a timely and cost-effective screening prior to considering whole exome sequencing as routine diagnostic tool. The use of targeted gene panels can shorten the course from seizure onset to genetic diagnosis. Current therapy for epileptic encephalopathies has mainly been focused on reducing the occurrence of seizure rather than influencing the underlying disease process. Based on limited genotype-phenotype correlations and available clinical evidences, some precision medicine in epilepsy has been proposed [42,43]. Improvements in comprehensive understanding of molecular pathogenesis underlying epileptic encephalopathies will pave the path to the development of effective precise therapeutic strategies for the diseases as early as possible.
Acknowledgement
The authors are grateful to Kevin Liu B.Sc. for administrative and technical supports and to Sofiva Genomics, Taipei, and Taiwan R.O.C. for genotyping assistance. The present study was funded by Far Eastern Memorial Hospital (99-FEMH-C-034, 100-FEMH-C-018, 100FJU-FEMH-04 and 103-FEMH-FJU-03), Taiwan, R.O.C.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Dulac O. Epileptic encephalopathy. Epilepsia 42(3), 23-26 (2001).
- McTague A, Howell KB, Cross JH, et al. The genetic landscape of the epileptic encephalopathies of infancy and childhood. Lancet. Neurol 15(3), 304-316 (2016).
- Guerrini R. Epilepsy in children. Lancet 367(9509), 499-524 (2006).
- Hwang SK, Kwon S. Early-onset epileptic encephalopathies and the diagnostic approach to underlying causes. Korean. J. Pediatr 58(11), 407-414 (2015).
- Nabbout R, Dulac O. Epileptic encephalopathies: a brief overview. J. Clin. Neurophysiol 20(6), 393-397 (2003).
- Lemke JR, Riesch E, Scheurenbrand T, et al. Targeted next generation sequencing as a diagnostic tool in epileptic disorders. Epilepsia 53(8), 1387-1398 (2012).
- Mercimek-Mahmutoglu S, Patel J, Cordeiro D, et al. Diagnostic yield of genetic testing in epileptic encephalopathy in childhood. Epilepsia 56(5), 707-716 (2015).
- Nieh SE, Sherr EH. Epileptic encephalopathies: new genes and new pathways. Neurotherapeutics 11(4), 796-806 (2014).
- Li H, Durbin R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26(5), 589-595 (2010).
- McKenna A, Hanna M, Banks E, et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome. Res 20(9), 1297-1303 (2010).
- Zupanc ML. Clinical evaluation and diagnosis of severe epilepsy syndromes of early childhood. J. Child. Neurol 24(8), 6S-14S (2009).
- Butler KM, da Silva C, Alexander JJ, et al. Diagnostic Yield From 339 Epilepsy Patients Screened on a Clinical Gene Panel. Pediatr. Neurol 77, 61-66 (2017).
- Trump N, McTague A, Brittain H, et al. Improving diagnosis and broadening the phenotypes in early-onset seizure and severe developmental delay disorders through gene panel analysis. J. Med. Genet 53(5), 310-317 (2016).
- Steinlein OK. Genetic mechanisms that underlie epilepsy. Nat. Rev. Neurosci 5(5), 400-408 (2004).
- Orrico A, Galli L, Grosso S, et al. Mutational analysis of the SCN1A, SCN1B and GABRG2 genes in 150 Italian patients with idiopathic childhood epilepsies. Clin. Genet 75(6), 579-581 (2009).
- Harkin LA, McMahon JM, Iona X, et al. The spectrum of SCN1A-related infantile epileptic encephalopathies. Brain 130(3), 843-852 (2007).
- Berkovic SF, Heron SE, Giordano L, et al. Benign familial neonatal-infantile seizures: characterization of a new sodium channelopathy. Ann. Neurol 55(4), 550-557 (2004).
- Howell KB, McMahon JM, Carvill GL, et al. SCN2A encephalopathy: A major cause of epilepsy of infancy with migrating focal seizures. Neurology 85(11), 958-966 (2015).
- Nakamura K, Kato M, Osaka H, et al. Clinical spectrum of SCN2A mutations expanding to Ohtahara syndrome. Neurology 81(11), 992-998 (2013).
- Liao Y, Anttonen AK, Liukkonen E, et al. SCN2A mutation associated with neonatal epilepsy, late-onset episodic ataxia, myoclonus, and pain. Neurology 75(16), 1454-1458 (2010).
- Sanders SJ, Murtha MT, Gupta AR, et al. De novo mutations revealed by whole-exome sequencing are strongly associated with autism. Nature 485(7397), 237-241 (2012).
- Gardella E, Becker F, Moller RS, et al. Benign infantile seizures and paroxysmal dyskinesia caused by an SCN8A mutation. Ann. Neurol 79(3), 428-436 (2016).
- Veeramah KR, O'Brien JE, Meisler MH, et al. De novo pathogenic SCN8A mutation identified by whole-genome sequencing of a family quartet affected by infantile epileptic encephalopathy and SUDEP. Am. J. Hum. Genet 90(3), 502-510 (2012).
- Wagnon JL, Meisler MH. Recurrent and Non-Recurrent Mutations of SCN8A in Epileptic Encephalopathy. Front. Neurol 6, 104 (2015).
- Barcia G, Fleming MR, Deligniere A, et al. De novo gain-of-function KCNT1 channel mutations cause malignant migrating partial seizures of infancy. Nat. Genet 44(11), 1255-1259 (2012).
- Torkamani A, Bersell K, Jorge BS, et al. De novo KCNB1 mutations in epileptic encephalopathy. Ann. Neurol 76(4), 529-540 (2014).
- Lim CX, Ricos MG, Dibbens LM, et al. KCNT1 mutations in seizure disorders: the phenotypic spectrum and functional effects. J. Med. Genet 53(4), 217-225 (2016).
- Soldovieri MV, Miceli F, Taglialatela M. Driving with no brakes: molecular pathophysiology of Kv7 potassium channels. Physiology (Bethesda) 26(5), 365-376 (2011).
- Nemos C, Lambert L, Giuliano F, et al. Mutational spectrum of CDKL5 in early-onset encephalopathies: a study of a large collection of French patients and review of the literature. Clin. Genet 76(4), 357-371 (2009).
- White R, Ho G, Schmidt S, et al. Cyclin-dependent kinase-like 5 (CDKL5) mutation screening in Rett syndrome and related disorders. Twin. Res. Hum. Genet 13(2), 168-178 (2010).
- Bahi-Buisson N, Villeneuve N, Caietta E, et al. Recurrent mutations in the CDKL5 gene: genotype-phenotype relationships. Am. J. Med. Genet A 158A(7), 1612-1619 (2012).
- Saitsu H, Kato M, Mizuguchi T, et al. De novo mutations in the gene encoding STXBP1 (MUNC18-1) cause early infantile epileptic encephalopathy. Nat. Genet 40(6), 782-788 (2008).
- Gursoy S, Ercal D. Diagnostic Approach to Genetic Causes of Early-Onset Epileptic Encephalopathy. J. Child. Neurol 31(4), 523-532 (2016).
- Yamatogi Y, Ohtahara S. Early-infantile epileptic encephalopathy with suppression-bursts, Ohtahara syndrome; its overview referring to our 16 cases. Brain. Dev 24(1), 13-23 (2002).
- Rickman C, Medine CN, Bergmann A, et al. Functionally and spatially distinct modes of munc18-syntaxin 1 interaction. J. Biol. Chem 282(16), 12097-12103 (2007).
- Kim JH, Lee HK, Takamiya K, et al. The role of synaptic GTPase-activating protein in neuronal development and synaptic plasticity. J. Neurosci 23(4), 1119-1124 (2003).
- Mignot C, von Stulpnagel C, Nava C, et al. Genetic and neurodevelopmental spectrum of SYNGAP1-associated intellectual disability and epilepsy. J. Med. Genet 53(8), 511-522 (2016).
- von Stulpnagel C, Funke C, Haberl C, et al. SYNGAP1 Mutation in Focal and Generalized Epilepsy: A Literature Overview and A Case Report with Special Aspects of the EEG. Neuropediatrics 46(4), 287-291 (2015).
- Liang JS, Lin LJ, Lin HC, et al. An unusual GFAP mutation in a Taiwanese child with infantile Alexander disease. Pediatr. Neonatol (2018).
- Brenner M, Johnson AB, Boespflug-Tanguy O, et al. Mutations in GFAP, encoding glial fibrillary acidic protein, are associated with Alexander disease. Nat. Genet 27(1), 117-120 (2001).
- Gorospe JR, Maletkovic J. Alexander disease and megalencephalic leukoencephalopathy with subcortical cysts: leukodystrophies arising from astrocyte dysfunction. Ment. Retard. Dev Disabil. Res. Rev 12(2), 113-122 (2006).
- Liang JS, Lin LJ, Yang MT, et al. The therapeutic implication of a novel SCN2A mutation associated early-onset epileptic encephalopathy with Rett-like features. Brain. Dev 39(10), 877-881 (2017).
- Reif PS, Tsai MH, Helbig I, et al. Precision medicine in genetic epilepsies: break of dawn? Expert. Rev. Neurother 17(4), 381-392 (2017).