Research Article - Neuropsychiatry (2018) Volume 8, Issue 1
Associations between DNA methylation of Promoter Exon IX, Serum Protein and mRNA levels of Brain-derived Neurotrophic factor in Patients with Bipolar Mania
- *Corresponding Author:
- Dr. Tiao-Lai Huang
Department of Psychiatry
Kaohsiung Chang Gung Memorial Hospital 123
Ta-Pei Road, Niao-Sung District, Kaohsiung 833, Taiwan
Tel: 886-7-7317123 ext. 8753
Fax: 886-7-7326817
Abstract
Abstract
Background: Brain-derived neurotrophic factor (BDNF) could be as a molecular candidate for the development of bipolar disorder. We tried to investigate the relationships among BDNF promoter exon IX gene methylation, BDNF protein, and mRNA levels between patients with bipolar mania and healthy controls in peripheral blood.
Methods: Thirty-nine patients with bipolar mania and 39 healthy controls were recruited in this study. Psychiatric diagnoses were made according to DSM-IV criteria. Clinical severity of mania was assessed by the Young Mania Rating Scale, and only those with a score greater than 26 were recruited. After four weeks of medical treatment, 34 patients agreed to a follow-up evaluation.
Keywords
Bipolar disorder, Brain-Derived Neurotrophic Factor, DNA Methylation, Mood stabilizer
Introduction
Several studies have mentioned that the brainderived neurotropic factor (BDNF) could be a molecular candidate for the development of bipolar disorder [1-8]. Previous studies had provided information on the relationship between serum BDNF protein levels and bipolar disorder [9-12]. In addition, the relationship between BDNF polymorphisms and bipolar disorder had also been discussed [4,12].
Recent studies have discussed epigenetic regulations in the human brain and peripheral blood [13-20]. Some of earlier studies focused on the epigenetic alterations of BDNF and its tropomyosin-related kinase B (TrkB) receptor promoters [15,16,18-21], particularly in DNA methylation, which had been shown to be affected by aging and gender [22,23].
Epigenetic mechanisms have been applied to animal models of depression at the BDNF promoter [24-26], particularly in exon I, exon IV, and exon IX. However, no data exist on the DNA methylation of the BDNF promoter for an animal model of bipolar mania. Previous studies have reported methylation of the BDNF promoter in exon I and IV in the peripheral blood of patients with severe mental disorders, including schizophrenia [27,28], bipolar mania [27-29], and major depression [18-20,30].
More specifically, D’ Addario, et al. reported data on DNA methylation of the BDNF exon I promoter in the peripheral blood of patients with bipolar mania [29]. They observed a significant BDNF gene expression down regulation in bipolar II, but not in bipolar I patients compared with controls, and a consistent hypermethylation of the BDNF promoter region in bipolar II patients. Roth et al. observed the lasting epigenetic influence of early life adversity on the BDNF gene in an animal model, including the exon IV and exon IX promoter. However, data on CpG sites near rs6265 (also known as G196A or Val66Met polymorphisms) in the BDNF region in peripheral blood have not been reported, particularly in BDNF exon IX (coding exon) [31].
Taken together, in this study, we simultaneously detected the associations in DNA methylation of the BDNF promoter exon IX, and BDNF protein and mRNA levels in peripheral blood between patients with bipolar mania and healthy controls.
Materials and Methods
▪ Subjects
From August 2010 to July 2012, November 2012 to October 2013, and December 2014 to November 2015, patients with bipolar mania and healthy controls were evaluated according to the DSM-IV criteria using a semi-structured interview [32]. Healthy controls were evaluated with the Chinese Health Questionnaire-12 [33]. Approval was obtained from the ethics committee of the Institutional Review Board of Chang Gung Memorial Hospital. All participants had the ability to provide written informed consent. All assessments were conducted by the same senior psychiatrist. Patients with bipolar mania who scored over 26 on the Young Mania Rating Scale (YMRS) [34] were recruited into this study. The severity of YMRS was assessed again after four weeks of medical treatment.
Mood stabilizers of either lithium (dose range, 900-1200 mg/d) or valproaic acid (dose range, 600–1500 mg/d) were prescribed during hospitalization. The antipsychotic drugs including risperidone (1–4 mg/d), olanzapine (5–15 mg/d), or clozapine (50-300 mg/d), as well as other psychotropic agents (i.e., lorazepam, 0–3 mg/d) were also prescribed along with the aforementioned mood stabilizers. The patients had not taken any drugs for at least 2 weeks prior to the recruitment of the study, nor were they heavy smokers or alcohol dependent. No participants exhibited systemic diseases, such as cardiovascular disease, liver disease, or thyroid disease.
▪ Assessing BDNF protein, mRNA, and DNA methylation
To analyze BDNF protein, mRNA, and DNA methylation levels in all participants at baseline, 15 mL of venous blood was obtained from each participant. Patients with bipolar mania were examined for biological markers after four weeks of medical treatment. Some of the data on BDNF protein and mRNA levels of some patients had previously been published in a preliminary study [35] as well as a recently accepted article [36].
▪ DNA isolation and bisulfite treatment
Genomic DNA was isolated using an Easy Blood Genomic DNA Purification Kit (GeneMark) according to the manufacturer instructions. Genomic DNA (500 ng) from lymphocytes was converted with sodium bisulfite and the EZ DNA Methylation Kit (Zymo Research) [37,38]. The concentration of sodium-bisulfatetreated DNA was measured using an ND-1000 spectrophotometer (NanoDrop Technologies), and 20 ng of treated DNA was used in a regionspecific PCR.
▪ Pyrosequencing analysis
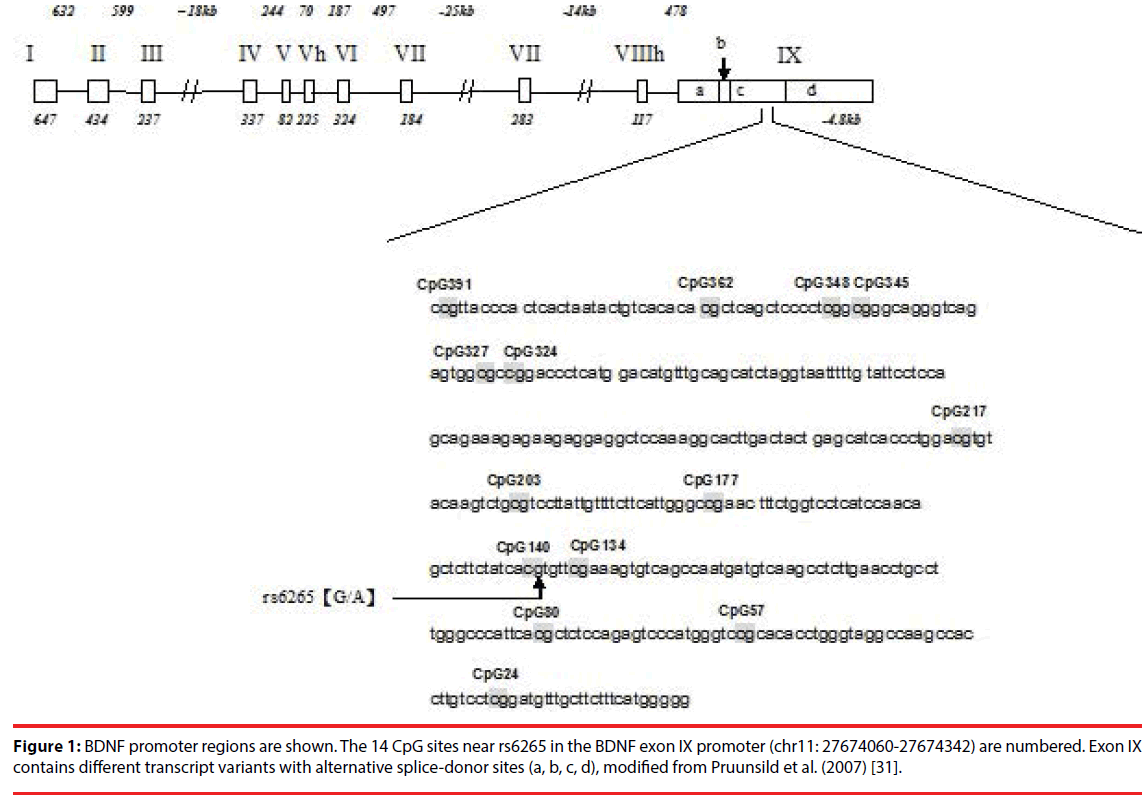
Pyrosequencing analysis was conducted for 14 CpG sites near rs6265 in the BDNF exon IX, using a PyroMark Q24 System (Biotage, Sweden) [31,39]. PCR was performed with 20 ng of bisulfite-treated DNA, 200 mM of each primer, 12.5 μL of PyroMark 2X PCR master mix, and 2.5 μL of CoralLoad Concentrate 10X (provided in the PyroMark PCR Master Mix, Qiagen) (Figure 1). Twenty microliters of DNA of each sample were used as the template in PCR reactions, using the following primers: BDNF_Forward 5’- TTT GGT TGT ATG AAG GTT GTT TTT ATG AA-3’; BDNF _Reverse 5’- ACA TAT CCA CTA CAA TCT TTT TAT CTA C-3’(5’ label biotin). PCR conditions were 95°C for 15 minutes; 45 cycles at 94°C for 30 seconds, 55°C for 30 seconds, 72°C for 40 seconds; and 72°C for 10 minutes. PCR products of 442 bp were examined using 1.5% agarose gel electrophoresis. The sequencing primer was as follows: BDNF_S1 5’-GGT TGT TTT TAT GAA AGA AGT-3’; BDNF_S2 5’- AGA GGT TTG ATA TTA TTG G-3’; and BDNF_S3 5’- GTT GTA AAT ATG TTT ATG AGG G-3’.
Figure 1: BDNF promoter regions are shown. The 14 CpG sites near rs6265 in the BDNF exon IX promoter (chr11: 27674060-27674342) are numbered. Exon IX contains different transcript variants with alternative splice-donor sites (a, b, c, d), modified from Pruunsild et al. (2007) [31].
Subsequent quantification of the methylation density of the 14 CpG sites was performed using PyroMark Q24 software.
▪ Serum BDNF protein levels
Serum BDNF protein levels were measured using a commercially available ELISA kit of the sandwich type (BDNF Emax Immunoassay System; Promega; USA). Each system contained anti-BDNF mAb, BDNF standard, antihuman BDNF pAb, Block&Sample 5X buffer, anti-IgY HRP, TMB solution, peroxidase substrate, and protocol. All samples were assayed and duplicated by the senior technologist.
▪mRNA isolation and reverse transcription polymerase chain reaction
Peripheral blood was collected using a PAXgene Blood RNA Tube (Qiagen) and extracted using a PAXgene Blood RNA Kit. The total RNA (1 microg) was reverse-transcribed into cDNA by using a High Capacity cDNA Reverse Transcription Kit (Applied Biosystems). The PCR of BDNF and beta-actin gene expression (with primer pairs ordered from Promega Biosciences) were conducted using SYBR Green (Applied Biosystems), using 5 microL of cDNA in a 20-microL final volume and 0.5 microM of each primer (final concentration). BDNF mRNA primers are Forward 5’- AAG AAG CAA ACA TCC GAG G -3’ and Reverse 5’- AAG GCA CTT GAC TAC TGA G -3’. Quantitative PCR was performed using a 7500 Fast Real-Time PCR System (Applied Biosystems) for 40 cycles at 95°C for 5 seconds, at a specific annealing temperature of 60°C for 5 seconds and 72°C for 12 seconds. Amplification specificity was investigated using the melting curve following the manufacturer instructions. The results were analyzed using 7500 Fast Real-Time PCR System Software v1.4.1 (Applied Biosystems). Gene expression levels were expressed as the concentration ratios between PCR products and beta-actin in the same sample.
▪ Data analysis
All results were presented as mean ± standard deviation. The statistical differences of BDNF protein levels, mRNA levels, and degrees of methylation of patients with bipolar mania and the healthy controls were determined through t tests or an analysis of covariance (ANCOVA) adjusted for age and gender. Pearson’s correlation was performed between DNA methylation at different CpG sites and disease severity (YMRS score), serum protein, and mRNA levels. A p value less than .05 was considered to be statistically significant.
Results
During study periods, 39 patients with bipolar mania and 39 healthy controls were recruited. Table 1 shows the demographic data as well as serum BDNF protein and mRNA levels at baseline of all participants. Among the 39 patients with bipolar mania, 34 agreed to be reevaluated after 4 weeks of medical treatment. Table 2 shows the demographic data as well as serum BDNF protein and BDNF mRNA levels at baseline and at endpoint after a 4-week treatment in patients with bipolar mania. Some of the data on BDNF protein and mRNA levels of some patients had previously been published in a preliminary study [35] as well as a recently accepted article [36].
| Diagnostic groups | Age | BMI (kg/m2) |
Education (years) |
Duration of illness (years) |
BDNF protein levels (ng/ml) |
BDNF mRNA levels |
YMRS |
|---|---|---|---|---|---|---|---|
| Bipolar mania (n=39) |
39.2 ± 11.8 | 24.3 ± 5.1 | 12.5 ± 1.7 | 13.3 ± 10.1 | 4.9 ± 3.8 | 1.0 ± 0.8 | 35.0 ± 9.9 |
| Men (n=18) | 44.0 ± 11.0 | 24.5 ± 5.1 | 12.8 ± 1.4 | 18.1 ± 8.6 | 4.9 ± 4.1 | 0.9 ± 1.0 | 33.0 ± 11.0 |
| Women (n=21) | 35.0 ± 11.0 | 24.1 ± 5.1 | 12.2 ± 2.0 | 9.2 ± 9.9 | 4.9 ± 3.7 | 1.1 ± 0.6 | 36.7 ± 8.8 |
| Healthy Controls (n=39) | 31.7 ± 6.0 | 22.4 ± 3.8 | 17.4 ± 1.8 | - | 9.5 ± 3.3 | 3.3 ± 3.4 | |
| Men (n=18) | 29.9 ± 4.7 | 23.8 ± 3.4 | 17.8 ± 1.7 | - | 8.4 ± 3.3 | 1.9 ± 2.2 | |
| Women (n=21) | 33.2 ± 6.6 | 21.3 ± 3.0 | 17.1 ± 1.8 | - | 10.4 ± 3.1 | 4.5 ± 3.9 |
BMI= body mass index; YMRS=Young Mania Rating Scale Score
Table 1: Demographic data and serum BDNF protein and BDNF mRNA levels at baseline in all participants.
| Age | Duration of illness (years) |
BDNF protein (baseline) ng/ml |
BDNF protein (endpoint) ng/ml |
BDNF mRNA (baseline) |
BDNF mRNA (endpoint) |
YMRS (baseline) |
YMRS (endpoint) |
|
|---|---|---|---|---|---|---|---|---|
| Total (n=34) |
38.4 ± 12.1 | 13.6 ± 9.9 | 4.7 ± 3.4 | 6.1 ± 5.9 | 0.9 ± 0.7 | 1.3 ± 1.8 | 35.1 ± 10.2 | 5.6 ± 7.7 |
| Men (n=16) |
43.6 ± 11.6 | 17.5 ± 8.2 | 4.5 ± 3.4 | 5.2 ± 6.8 | 0.7 ± 0.7 | 1.3 ± 1.6 | 32.3 ± 10.8 | 4.3 ± 3.8 |
| Women (n=18) |
33.8 ± 10.8 | 10.1 ± 10.2 | 4.9 ± 3.6 | 6.9 ± 5.0 | 1.0 ± 0.6 | 1.4 ± 1.9 | 37.5 ± 9.3 | 6.7 ± 9.9 |
The values are presented as mean ± standard deviation.
BDNF: brain-derived neurotrophic factor; YMRS: Young Mania Rating Scale.
Table 2: Demographic data and serum BDNF protein and BDNF mRNA levels at baseline and at endpoint of a 4-week treatment in all participants.
▪ BDNF promoter exon IX gene methylation sequential analysis
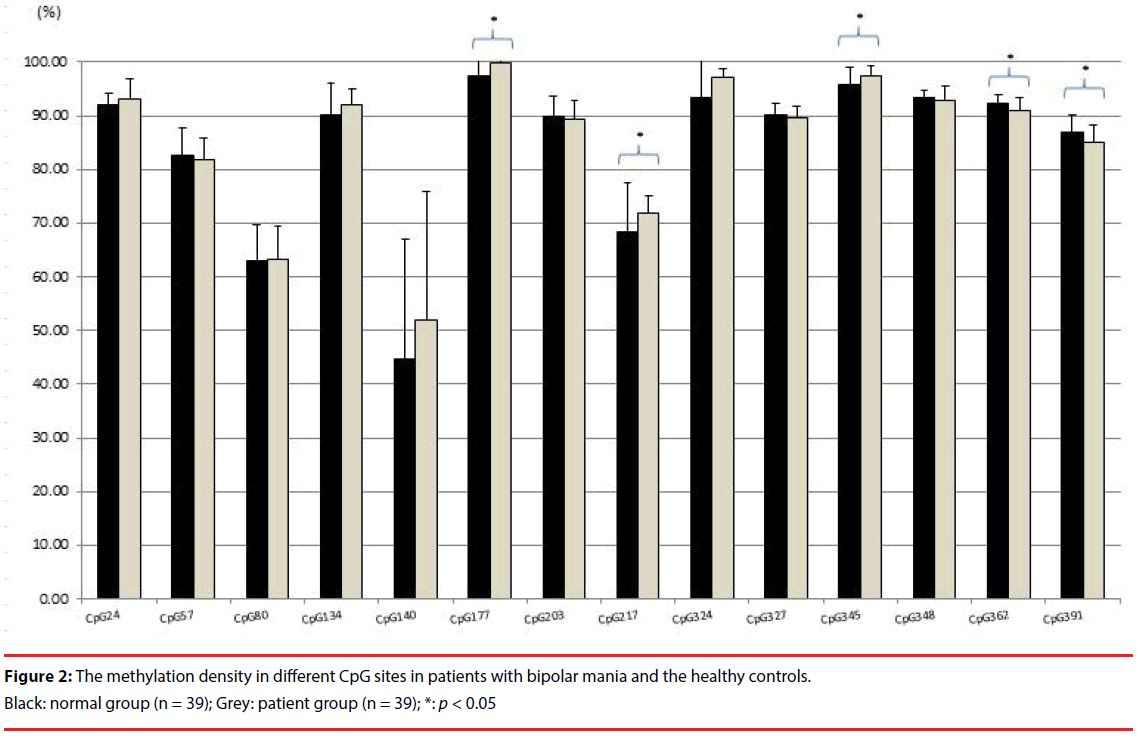
The BDNF promoter exon IX gene methylation sequential analysis was completed for the 39 patients with bipolar mania and 39 healthy controls at baseline. Figure 2 shows the methylation density of investigated CpG sites between patients with bipolar mania and healthy controls at baseline. Using t-test, patients with bipolar mania had higher degrees of methylation at CpG sites 177 (t = 2.782, p = .007) and 217 (t = 2.274, p = .026), and a lower degree of methylation at CpG sites 345 (t = - 2.660, p = .010), 362 (t = -3.142, p = .002), and 391 (t = -2.747, p = .008), than did the healthy controls. Using ANCOVA adjusted for age and gender, statistical significance remained at CpG sites 217 (p < .001) and 391 (p = 0.013).
Pearson’s correlation found no statistically significant association between YMRS score and degrees of methylation at studied CpG sites. Serum BDNF protein level correlated significantly with the degree of methylation at CpG site 348 (r=.290, p = .010), but not with other CpG sites. BDNF mRNA level correlated significantly with degree of methylation at CpG sites 134 (r=-.319, p = .004), 177 (r=-.490, p < .001) and 217 (r=-.460, p < .001).
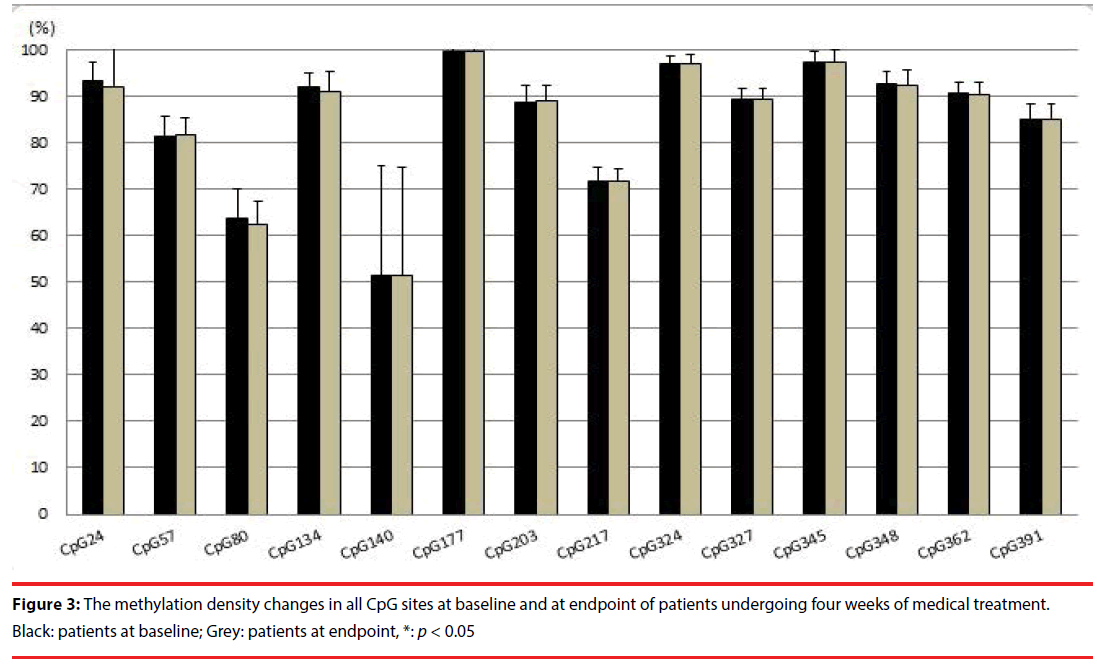
After four weeks of medical treatment, using paired t test, degrees of methylation of fourteen studied CpG sites did not change significantly in the 34 patients with bipolar mania (Figure 3). Using paired t-test, YMRS scores decreased significantly after treatment (p < 0.001). At baseline, YMRS score did not correlate with the degree of methylation of the studied CpG sites in the blood of the 34 patients with bipolar mania. After treatment, YMRS score significantly correlated with the degree of methylation at CpG site 391 (r = -0.441, p = 0.009), but not with the others.
Serum BDNF protein level
Using t test, at baseline, patients with bipolar mania had lower BDNF protein levels than did the healthy controls (t = 5.685, p < .001). Using ANCOVA adjusted for age and gender,the statistical significance was lost (F = 1.872, p = .176).
Using paired t test, after four weeks of medical treatment, serum BDNF protein levels showed no significant change (t = 1.374, p = .179).
▪ Blood BDNF mRNA
Using t test, at baseline, patients with bipolar mania had lower BDNF mRNA levels than did the healthy controls (t = -4.100, p < .001). Using ANCOVA adjusted for age and gender, the statistical significance persisted (F = 7.561, p = .008).
Using paired t test, after four weeks of medical treatment, serum BDNF protein levels showed no significant change (t = 1.252, p = .219).
Discussion
The first major finding of this study is that patients with bipolar mania had a higher degree of methylation at CpG site 217 (p < .001) and lower degree of methylation at CpG site 391 (p = 0.13) than did the healthy controls in BDNF exon IX, though no difference was detected at CpG site 140 (also known as G196A, Val66Met, or rs6265 polymorphisms). While the methylation of BDNF gene had been studied in the past, data on exon IX was relatively scarce.
In the postmortem frontal cortex of 35 patients with schizophrenia and 35 patients with bipolar disorder, Val homozygotes of rs6265 had significantly higher DNA methylation across the exonic region profiled with pyrosequencing [13]. We did not perform genotyping so a comparison could not be made. In the postmortem frontal cortex of ten patients with bipolar disorder, global DNA hypermethylation and BDNF CpG hypermethylation were found, though the paper did not specify the exons investigated [40]. A study investigating BDNF exon I found higher degree of methylation of BDNF and lower mRNA level in the blood of patients with bipolar II diorder, but not patients with bipolar I disorder [29]. That study also found that patients treated with both mood stabilizers and antidepressants had higher methylation levels than the patients treated with mood stabilizers alone, and that treatment with lithium or valproate was associated with significant reduction of methylation compared to other drugs. In our study we found no significant change in the degrees of methylation at 14 CpG sites of exon IX after four weeks of lithium and valproate treatment. This discrepancy could be caused by the difference in studied exons. In a study investigating methylation difference of BDNF promoters III and V in the leukocytes of 50 patients of bipolar I disorder and 50 age and sex matched controls, five CpG sites (one in III and five in V) out of 36 CpG sites showed significantly higher methylation levels [41]. In another study investigating BDNF exon I in the peripheral blood, higher methylation was found in patients with major depressive disorder than in either bipolar disorder or control group, and higher methylation was associated with antidepressant treatment [42]. In a follow-up of D’Addario et al’s work focusing on BDNF exon I in peripheral blood samples, higher methylation was found again in patients with bipolar II diosrder as compared to patients with bipolar I disorder or major depressive disorder, and higher methylation levels was found in patients with depressed state as compared to patients with manic or mixed state [43]. That study continued to find that treatment with lithium or valproate was associated with lower methylation levels in BDNF exon I. Overall, the investigated regions of BDNF methylation could vary widely between the studies, making direct comparisons challenging, not to mention the methodological and analytical differences [44]. Nevertheless, the various findings could help a better understanding of underlying mechanism of bipolar disorder.
We also found that serum BDNF protein level correlated significantly with the degree of methylation at CpG site 348, and that BDNF mRNA level correlated significantly with degree of methylation at CpG sites 134, 177, and 217. After four weeks of medical treatment, YMRS score significantly correlated with the degree of methylation at CpG site 391 (r = -0.441, p = 0.009). Whether the degrees of methylation of those CpG sites could be used to infer the levels of BDNF protein and mRNA levels or predict the treatment response will require a larger sample size to verify.
In addition, our data simultaneously showed that patients with bipolar mania exhibited lower BDNF protein and mRNA levels than did the healthy controls, which is compatible with certain reports [10,34,37,38], but not with others [45,46].
There are several limitations to this study. The sample size was relatively small. The investigated region of BDNF was limited. No genotyping was performed. The treatment period was only four weeks. The combined use of mood stabilizer and antipsychotics for managing bipolar mania could also have an effect on the DNA methylation.
Conclusion
These results indicated that the degree of BDNF promoter exon IX gene methylation in peripheral blood might be involved in the psychopathology bipolar mania and its treatment response. However, a large sample is required to verify those results.
Acknowledgements
This study was supported by clinical research grants from Chang Gung Memorial Hospital (CMRPG8D1471 and CMRPG8B0761) and Ministry of Science and Technology (previously National Science Council) (NSC99-2628-B- 182-002-MY2), Taiwan. We did not receive any financial support from any pharmaceutical company. All authors declare no conflict of interest.
References
- Karege F, Schwald M, El Kouaissi R. Drug-induced decrease of protein kinase a activity reveals alteration in BDNF expression of bipolar affective disorder. Neuropsychopharmacology 29(4), 805-812 (2004).
- Karege F, Schwald M, Papadimitriou P, et al. The cAMP-dependent protein kinase A and brain-derived neurotrophic factor expression in lymphoblast cells of bipolar affective disorder. J. Affect. Disord 79(1-3), 187-192 (2004).
- Frey BN, Andreazza AC, Ceresér KM, et al. Effects of mood stabilizers on hippocampus BDNF levels in an animal model of mania. Life. Sci 79(3), 281-286 (2006).
- Okada T, Hashimoto R, Numakawa T, et al. A complex polymorphic region in the brain-derived neurotrophic factor (BDNF) gene confers susceptibility to bipolar disorder and affects transcriptional activity. Mol. Psychiatry 11(7), 695-703 (2006).
- Tsai SJ. TrkB partial agonists: potential treatment strategy for epilepsy, mania, and autism. Med. Hypotheses 66(1), 173-175 (2006).
- Post RM. Role of BDNF in bipolar and unipolar disorder: clinical and theoretical implications. J. Psychiatr. Res 41(12), 979-990 (2007).
- Kapczinski, F., et al., Brain-derived neurotrophic factor and neuroplasticity in bipolar disorder. Expert Rev Neurother, 2008. 8(7): p. 1101-13.
- Stertz L, Fries GR, Aguiar BW, et al. Histone deacetylase activity and brain-derived neurotrophic factor (BDNF) levels in a pharmacological model of mania. Rev. Bras. Psiquiatr 36(1), 39-46 (2013).
- Lin PY. State-dependent decrease in levels of brain-derived neurotrophic factor in bipolar disorder: a meta-analytic study. Neurosci. Lett 466(3), 139-143 (2009).
- Tramontina JF, Andreazza AC, Kauer-Sant'anna M, et al. Brain-derived neurotrophic factor serum levels before and after treatment for acute mania. Neurosci. Lett 452(2), 111-113 (2009).
- Huang TL, Hung YY, Lee CT, et al. Serum protein levels of brain-derived neurotrophic factor and tropomyosin-related kinase B in bipolar disorder: effects of mood stabilizers. Neuropsychobiology 65(2), 65-69 (2012).
- Chen SL, Lee SY, Chang YH, et al. The BDNF Val66Met polymorphism and plasma brain-derived neurotrophic factor levels in Han Chinese patients with bipolar disorder and schizophrenia. Prog. Neuropsychopharmacol. Biol. Psychiatry 51(1), 99-104 (2014).
- Mill J, Tang T, Kaminsky Z, et al. Epigenomic profiling reveals DNA-methylation changes associated with major psychosis. Am. J. Hum. Genet 82(3), 696-711 (2008).
- Akbarian S, Huang HS. Epigenetic regulation in human brain-focus on histone lysine methylation. Biol. Psychiatry 65(3), 198-203 (2009).
- Ernst C, Deleva V, Deng X, et al. Alternative splicing, methylation state, and expression profile of tropomyosin-related kinase B in the frontal cortex of suicide completers. Arch. Gen. Psychiatry 66(1), 22-32 (2009).
- Mellios N, Huang HS, Baker SP, et al. Molecular determinants of dysregulated GABAergic gene expression in the prefrontal cortex of subjects with schizophrenia. Biol. Psychiatry 65(12), 1006-1014 (2009).
- Sweatt JD. Experience-dependent epigenetic modifications in the central nervous system. Biol. Psychiatry 65(3), 191-197 (2009).
- Fuchikami M, Morinobu S, Segawa M, et al. DNA methylation profiles of the brain-derived neurotrophic factor (BDNF) gene as a potent diagnostic biomarker in major depression. PLoS. One 6(8), e23881 (2011).
- Kang HJ, Kim JM, Lee JY, et al. BDNF promoter methylation and suicidal behavior in depressive patients. J. Affect. Disord 151(2), 679-685 (2013).
- Tadic A, Müller-Engling L, Schlicht KF, et al. Methylation of the promoter of brain-derived neurotrophic factor exon IV and antidepressant response in major depression. Mol. Psychiatry 19(3), 281-283 (2014).
- Dwivedi Y, Rizavi HS, Conley RR, et al. Altered gene expression of brain-derived neurotrophic factor and receptor tyrosine kinase B in postmortem brain of suicide subjects. Arch. Gen. Psychiatry 60(8), 804-815 (2003).
- Hannum G, Guinney J, Zhao L, et al. Genome-wide methylation profiles reveal quantitative views of human aging rates. Mol. Cell 49(2), 359-367 (2013).
- Boks MP, Derks EM, Weisenberger DJ, et al. The relationship of DNA methylation with age, gender and genotype in twins and healthy controls. PLoS. One 4(8), e6767 (2009).
- Tsankova NM, Berton O, Renthal W, et al. Sustained hippocampal chromatin regulation in a mouse model of depression and antidepressant action. Nat. Neurosci 9(4), 519-525 (2006).
- Tsankova N, Renthal W, Kumar A, et al. Epigenetic regulation in psychiatric disorders. Nat. Rev. Neurosci 8(5), 355-367 (2007).
- Roth TL, Lubin FD, Funk AJ, et al. Lasting epigenetic influence of early-life adversity on the BDNF gene. Biol. Psychiatry 65(9), 760-769 (2009).
- Roth TL, Lubin FD, Sodhi M, et al. Epigenetic mechanisms in schizophrenia. Biochim. Biophys. Acta 1790(9), 869-877 (2009).
- Ikegame T, Bundo M, Murata Y, et al. DNA methylation of the BDNF gene and its relevance to psychiatric disorders. J. Hum. Genet 58(7), 434-438 (2013).
- D'Addario C, Dell'Osso B, Palazzo MC, et al. Selective DNA methylation of BDNF promoter in bipolar disorder: differences among patients with BDI and BDII. Neuropsychopharmacology 37(7), 1647-1655 (2012).
- Keller S, Sarchiapone M, Zarrilli F, et al. Increased BDNF promoter methylation in the Wernicke area of suicide subjects. Arch. Gen. Psychiatry 67(3), 258-267 (2010).
- Pruunsild P, Kazantseva A, Aid T, et al. Dissecting the human BDNF locus: bidirectional transcription, complex splicing, and multiple promoters. Genomics 90(3), 397-406 (2007).
- American Psychiatric Association, Diagnostic and Statistical Manual of Mental Disorders Fourth Edition (Text Revision). Washington DC: American Psychiatric Publishing, Inc (2000).
- Chong MY, Wilkinson G. Validation of 30- and 12-item versions of the Chinese Health Questionnaire (CHQ) in patients admitted for general health screening. Psychol. Med 19(2), 495-505 (1989).
- Young RC, Biggs JT, Ziegler VE, et al. A rating scale for mania: reliability, validity and sensitivity. Br. J. Psychiatry 133(1), 429-435 (1978).
- Lin CC, Lee CT, Lo YT, et al. Brain-derived neurotrophic factor protein and mRNA levels in patients with bipolar mania - A preliminary study. Biomed. J 39(4), 272-276 (2016).
- Wu YS, Chiou YJ, Huang TL. Associations between Serum Brain-Derived Neurotrophic Factors and Bipolar Disorder. Neuropsychiatry 7(7) (2017).
- Bibikova M, Lin Z, Zhou L, et al. High-throughput DNA methylation profiling using universal bead arrays. Genome. Res 16(3), 383-393 (2006).
- Tai KY, Shiah SG, Shieh YS, et al. DNA methylation and histone modification regulate silencing of epithelial cell adhesion molecule for tumor invasion and progression. Oncogene 26(27), 3989-3997 (2007).
- Tsiatis AC, Norris-Kirby A, Rich RG, et al. Comparison of Sanger sequencing, pyrosequencing, and melting curve analysis for the detection of KRAS mutations: diagnostic and clinical implications. J. Mol. Diagn 12(4), 425-432 (2010).
- Rao JS, Keleshian VL, Klein S, et al. Epigenetic modifications in frontal cortex from Alzheimer's disease and bipolar disorder patients. Transl. Psychiatry 2(1), e132 (2012).
- Strauss JS, Khare T, De Luca V, et al. Quantitative leukocyte BDNF promoter methylation analysis in bipolar disorder. Int. J. Bipolar. Disord 1(1), 28 (2013).
- Carlberg L, Scheibelreiter J, Hassler MR, et al. Brain-derived neurotrophic factor (BDNF)-epigenetic regulation in unipolar and bipolar affective disorder. J. Affect. Disord 168(1), 399-406 (2014).
- Dell'Osso B, D'Addario C, Carlotta Palazzo M, et al. Epigenetic modulation of BDNF gene: differences in DNA methylation between unipolar and bipolar patients. J. Affect. Disord 166(1), 330-333 (2014).
- Mitchelmore C, Gede L. Brain Derived Neurotrophic Factor: epigenetic regulation in psychiatric disorders. Brain. Res 1586(1), 162-172 (2014).
- Dias VV, Brissos S, Frey BN, et al. Cognitive function and serum levels of brain-derived neurotrophic factor in patients with bipolar disorder. Bipolar. Disord 11(6), 663-671 (2009).
- Kauer-Sant'Anna M, Kapczinski F, Andreazza AC, et al. Brain-derived neurotrophic factor and inflammatory markers in patients with early- vs. late-stage bipolar disorder. Int. J Neuropsychopharmacol 12(4), 447-458 (2009).